News
Feed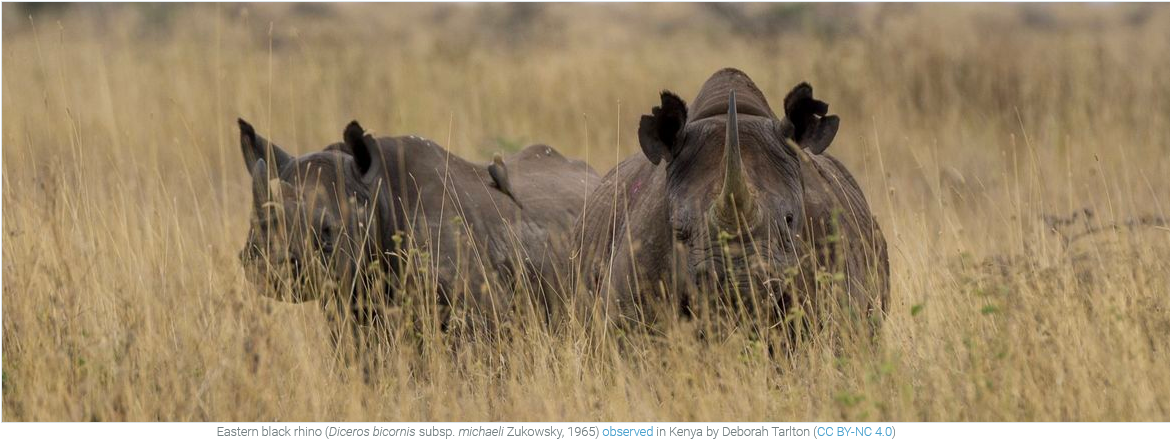
More than 10,000 scientific papers enabled by GBIF-mediated data
Significant growth in use of data from the GBIF network continues to demonstrate the importance and value of open biodiversity data

2024 Ebbe Nielsen Challenge seeks open-data innovations for biodiversity
The call for entries for the 2024 Ebbe Nielsen Challenge is now open!

GBIF to serve as administrative host for Species 2000 Secretariat
Agreement between GBIF Secretariat and the Catalogue of Life’s legal body will support further collaboration on ChecklistBank and other joint activities

SBDI Days 2024: Towards Data-driven Ecology
The SBDI consortium welcomes everyone within the scientific community who is curious about data-driven research. Join us at the Swedish Museum of Natural History in Stockholm.

Call opens for nominations to 2024 GBIF Graduate Researchers Award
Deadline for receiving nominations from GBIF member countries for graduate students whose innovative research relies on biodiversity data: 24 June 2024
GBIF Report Reveals Returns on Investments
The Global Biodiversity Information Facility (GBIF) has published a report shedding light on the returns yielded by investments in their data infrastructure. The report, titled “Unveiling the Value: The Return on Investments in GBIF,” highlights the wide-ranging impact of GBIF’s efforts on both scientific advancement and societal progress.

Guide on how to publish DNA-derived data on marine life
There is now an updated version of a guide to publishing sequence-based data. It is a practical guide for holders of genomic and metagenomic information, to include recommendations for sharing occurrence records based on DNA-based studies of the world’s oceans and seas.
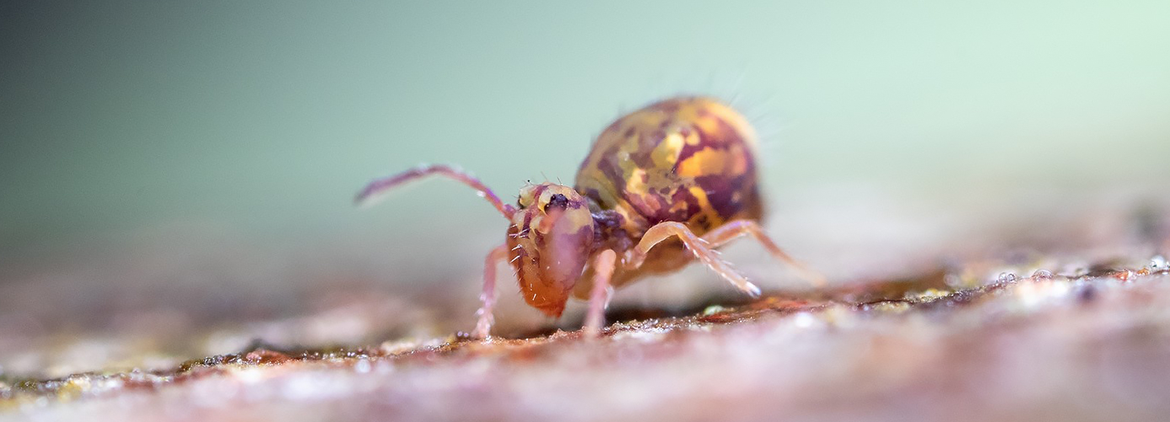

Share your dirt: Call for data papers describing datasets on soil biodiversity
GBIF continues partnership with FinBIF and Pensoft to support publication of new datasets about organisms whose life cycles depend on soil. DEADLINE: 22 September 2023

2023 Ebbe Nielsen Challenge seeks open-data innovations for biodiversity
Call for entries for submissions vying for €20,000 in prizes from annual incentive competition. DEADLINE: 22 August 2023




